There are two factors that play an integral role in distinguishing between populations within a species.
First, an organism’s geography restricts its ability to disperse. Individuals in adjacent populations migrate between one another at a higher rate than do individuals in more isolated populations. This leads to high gene flow between geographically close populations; the result of this is two groups that are difficult to genetically differentiate. In most ecosystems, the magnitude of differentiation increases with geographic distance.
Second, ecological and environmental differences can possibly cause genetic differences between populations within a species. In such environmentally heterogeneous conditions, interspecific differentiation (the difference between organisms of separate species) may increase with either geographic distance or ecological change. Questions arise then on whether ecological parameters like habitat type and climate play substantial roles in the formation of distinct populations or if populations differentiate due to geographic isolation.
Check out this video for a visual representation of these concepts!
Honey bees as model organisms

In order to understand which factor(s) cause genetic differences among a species, we first need to find a model organism having similar conditions in nature. Cue the honey bee: The honey bee (Arthropoda, Insecta, Hymenoptera) is a social eukaryotic animal which has 10 species and 26 subspecies distributed across the world, reaching all continents except Antarctica.
This cosmopolitan animal has adapted to many diverse ecotypes and already the complete genome of the European honey bee species Apis mellifera L. has been sequenced, meaning that we know what sequence of nucleotides makes up the entire DNA of the European honey bee. For these reasons, the honey bee has more recently been used as a model organism in population genetics studies.
Several honey bee subspecies (races) inhabiting South Africa are recognized as invasive races and have posed threats on African and U.S. beekeeping industries and agricultural agencies. In tropical Africa, there is significant geographical variability within honey bee races in spite of a lack of physical barriers (mountain ranges, large bodies of water, etc.). It is believed that the adaptation of honey bee races to a particular habitat is the basic mechanism of their genetic differentiation within a species. Now, several questions have arisen on how we can distinguish African honey bee races from one another. Do they have distinct genetic traits or do all races belong to the single population? What multidisciplinary approaches can we apply to identify these races? What is the magnitude of intra/interspecific genetic diversity and population structure? To get at these questions we first needed to collect a huge number of honey bees from South African colonies. With these specimens, we conducted an integrated investigation using genomic techniques at the Honey Bee Research and Extension Laboratory (HBREL), University of Florida.
What’s happening at HBREL?
To get at the above questions we first needed to collect a huge number of honey bees from South African colonies. With these specimens, we conducted an integrated investigation using genomic techniques at the Honey Bee Research and Extension Laboratory (HBREL), University of Florida.
In my current appointment as a postdoc in HBREL, I am leading several potential projects. Initially, I am focused on determining the genetic diversity and population structure as well as ultimately finding genetic markers among 900 individual bees (from 75 different apiaries and 29 geographical regions) of two unwanted honey bee races (Apis mellifera scutellata and Apis mellifera capensis) in South Africa. The primary purpose of this project is to establish genetic markers that can be used to identify potentially invasive honey bee species and subspecies, principally A. mellifera capensis Eschscholtz (the cape bee), and A.m. scutellata Lepeletier (the “African” or “killer” bee). This overall purpose benefits beekeepers directly because these bees are considered significant threats to the U.S. beekeeping industry. The proposed project offers an indirect benefit to the U.S. public since honey bees are the primary pollinators of many of the nation’s crops; any threat to honey bees is a threat to much of the U.S. food supply. It is also important to know that most invasive species jeopardize biodiversity and can have adverse effects on population dynamics of native bees.
Two South African honey bees are unwanted races threatening the African and U.S. honey bee populations in several primary ways; they may (1) out-compete the managed European-derived subspecies of honey bees in U.S., leading to lower profit margins for beekeepers, (2) present undesirable phenotypes, as was the case with A.m. scutellata, which is aggressively defensive, and (3) reduce the population of honey bees, due to social parasitic behavior of A.m. capensis.
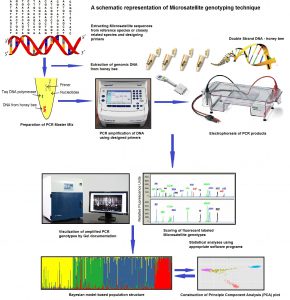
Developing advanced genetic markers would enable us to specifically identify honey bee populations with more efficient and precise strategies. As a first step in HBREL, we use a technique known as fluorescent labeled Microsatellite genotyping. Microsatellites are co-dominant markers with two to six nucleotide bases (nucleotide bases make up the structure of DNA), and are found across the nuclear genome of most organisms. These markers have rapidly become choice markers in population genetics studies due to their reproducibility and high rates of polymorphism (they occur in several different forms). Microsatellites are valuable tools for determining intra- and interspecific genetic variability and population genetic structure among honey bee populations. The outcomes of the present study will be presented at XXV International Congress of Entomology, September 25 – 30, in Orlando.
Have something to say? Join the discussion on our Facebook Page (www.facebook.com/UFHoneyBeeLab/).
 0
0
